Common in vitro methodologies for identifying the preferred DNA-binding sequences of transcription factors include systematic evolution of ligands by exponential enrichment (SELEX) accompanied with high-throughput sequencing (SELEX-seq) and high-density double-stranded DNA (dsDNA) microarrays ( 6). In vivo-based methods include chromatin immunoprecipitation (ChIP)-seq ( 4) and ChIP-chip ( 5), wherein a transcription factor of interest is immunoprecipitated from cross-linked biological samples and the resulting protein-bound DNA sequences are analyzed by high-throughput sequencing (ChIP-seq) or DNA microarray analysis (ChIP-chip). 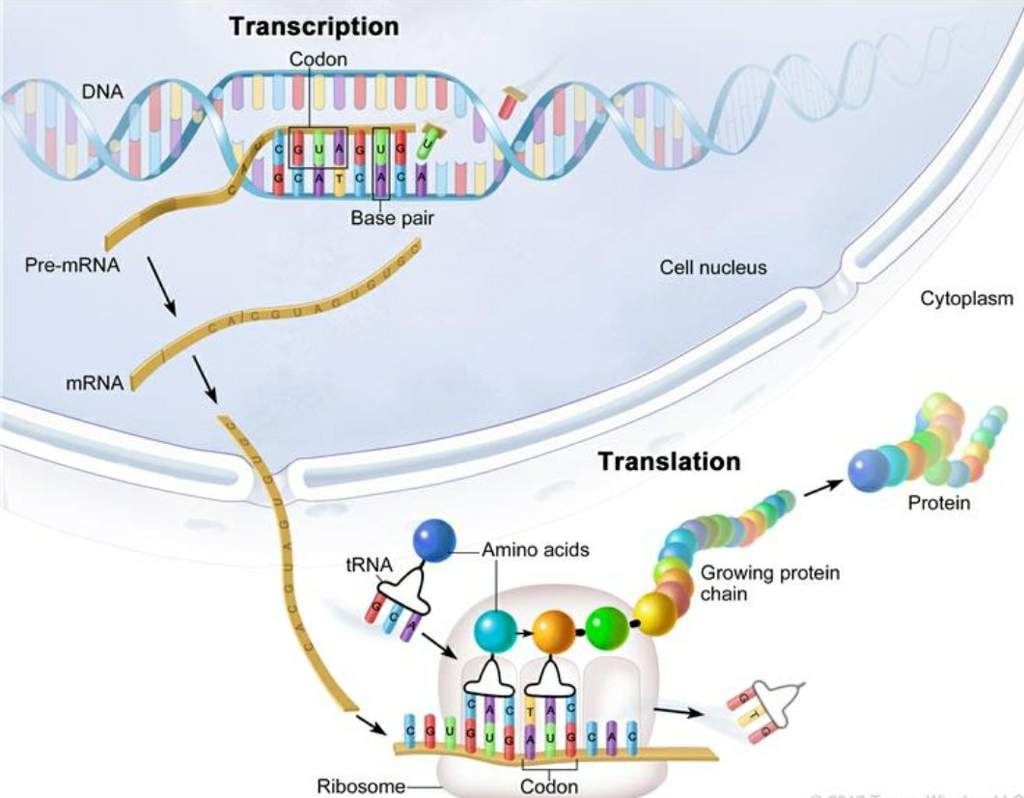
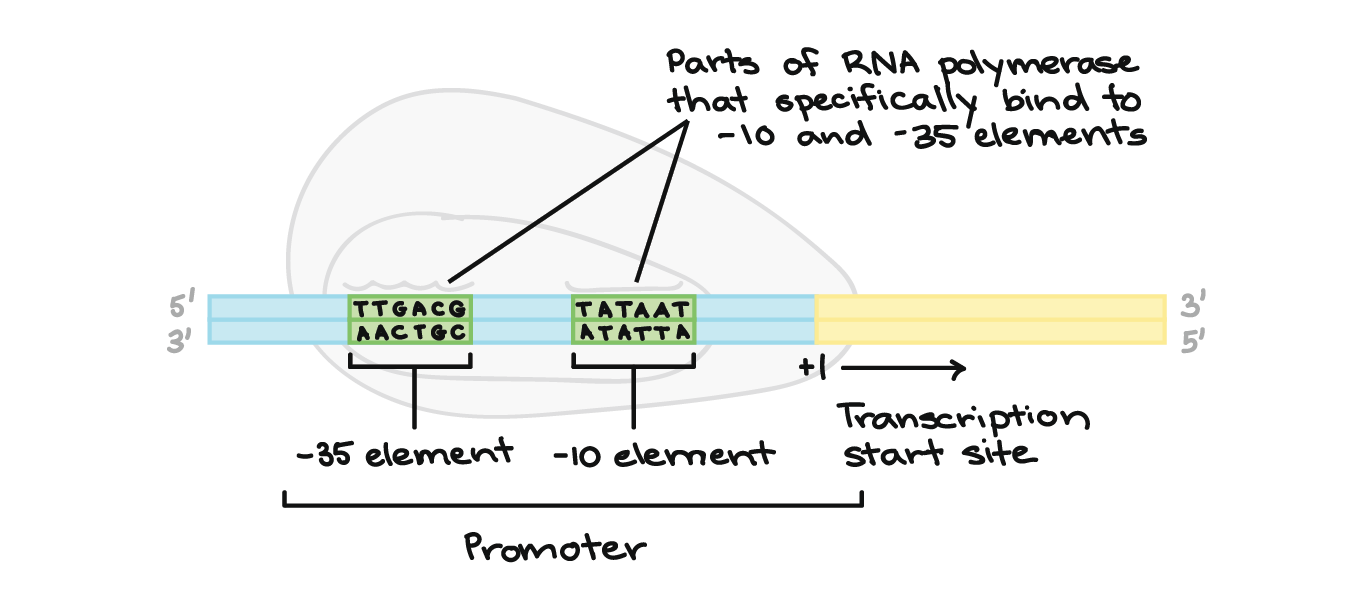
Indeed, several high-throughput experimental techniques have been developed to identify DNA-binding sequences of transcription factors in vivo and in vitro. Identifying the preferred DNA-binding sequence of a transcription factor presents an alternative starting strategy to discovering transcription regulatory networks, compared to transcriptome-wide analyses. As such, determination of transcription regulatory networks should, at minimum, include evidence of sequence-specific DNA binding and dysregulated gene expression in vivo in transcription factor knockout or overexpression strains. Many transcription factors, especially from prokaryotic organisms, recognize specific DNA motifs often found in the promoters of genes within their transcription regulatory network ( 3). These transcriptome-wide analyses yield hundreds of dysregulated genes, when only a small fraction are directly regulated by the transcription factor in question ( 2). Often, studies involving transcription regulatory network identification begin with differential gene expression analysis between a reference and transcription factor mutant strain using high-throughput means such as RNA sequencing or DNA microarrays. Extensive research has been conducted to identify transcription regulatory networks for transcription factors, which provide information on the transcription factor’s biological function ( 1). Transcription regulatory proteins, often referred to as transcription factors, are an essential class of proteins that help control gene expression, predominantly through modulating RNA polymerase activity. Cells have evolved various means to sense extracellular and intracellular stimuli and adapt their transcriptome accordingly. Regulation of gene expression is a fundamental biological mechanism for both prokaryotic and eukaryotic organisms. 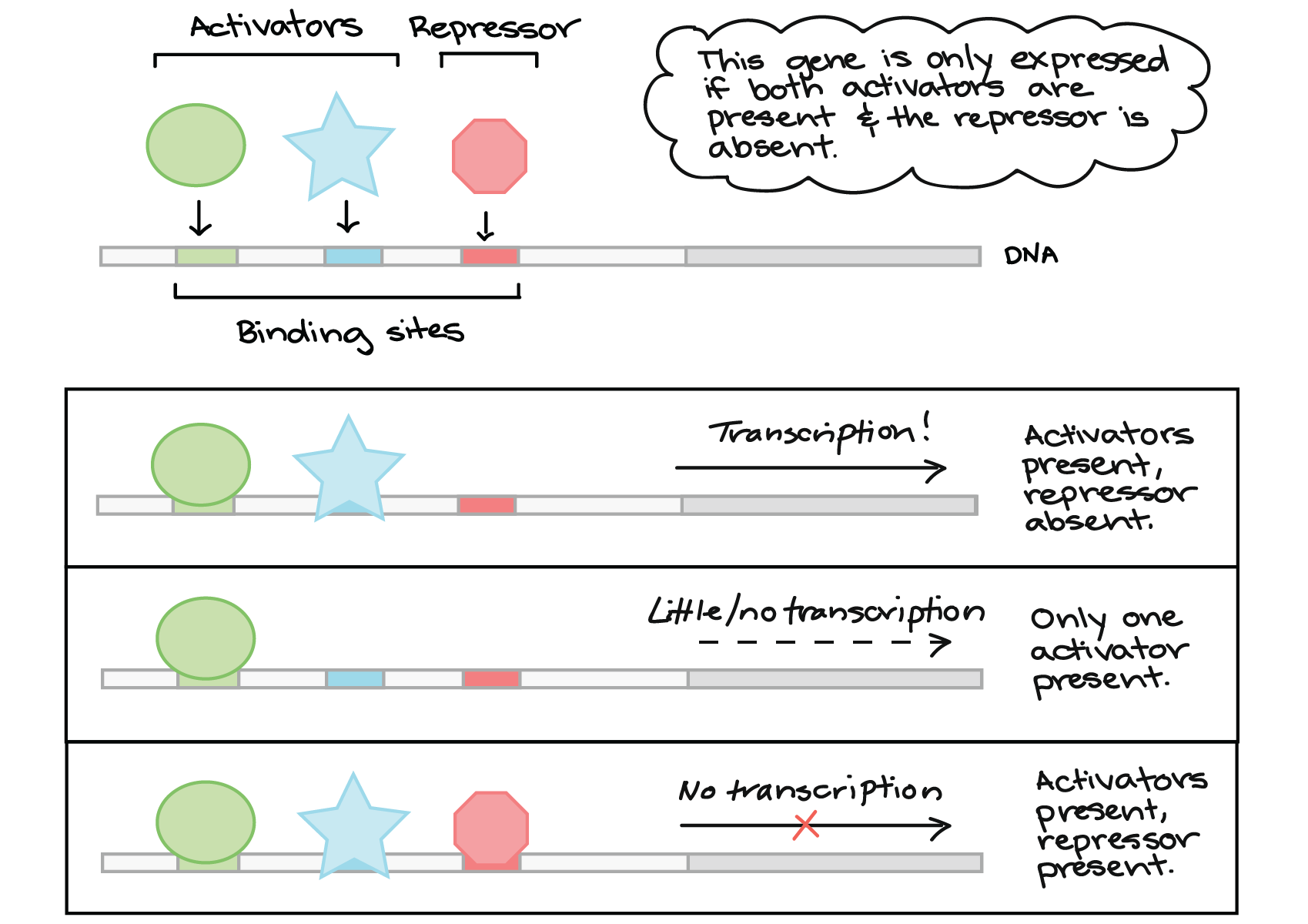
Subsequent high-throughput sequencing of REPSA-selected DNAs accompanied by motif discovery and scanning analyses can be used for determining transcription factor consensus binding sequences and potential regulated genes, providing critical first steps in determining organisms’ transcription regulatory networks. This process is repeated until a protected DNA species is observed by gel electrophoresis, which is an indication of a successful REPSA experiment. Cleavage-resistant DNA species are amplified by PCR and then used as inputs for the next round of REPSA. Contrary to traditional antibody-based capture methods, REPSA selects for transcription factor-bound DNA sequences by challenging binding reactions with a type IIS restriction endonuclease. In this protocol, we describe an alternative in vitro, iterative selection approach to ascertaining DNA-binding sequences of a transcription factor of interest using restriction endonuclease, protection, selection, and amplification (REPSA). Several assays, including SELEX-seq and ChIP-seq, capture DNA-bound transcription factors to determine the preferred DNA-binding sequences, which can then be mapped to the host organism’s genome to identify candidate regulatory genes. Various in vivo and in vitro assays have been developed to elucidate transcription regulatory networks. Research laboratories have devoted many projects to understanding the transcription regulatory networks for transcription factors, as these regulated genes provide critical insight into the biology of the host organism. Transcription factor proteins often bind regulatory DNA sequences upstream of transcription start sites to facilitate the activation or repression of RNA polymerase. Regulation of gene expression is a vital component of cellular biology. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
